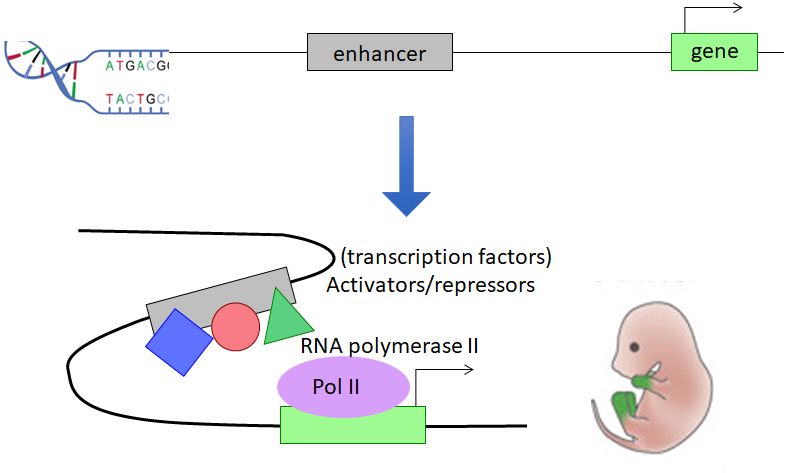
Enhancers
Enhancers are key regions of DNA that can be located upstream, downstream, even 1Mb away from a gene. Enhancers contain binding sites for proteins, and these proteins act as either activators or repressors, working together to promote or inhibit gene expression. However, gene regulation is not a static process. Genes are constantly being turned on and off depending on the current stage of development as well as location within an organism. It is the job of the enhancer to make sure genes are expressed at the right time and in the right part of the body.
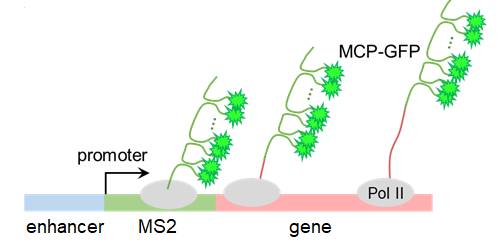
Quantitative Imaging
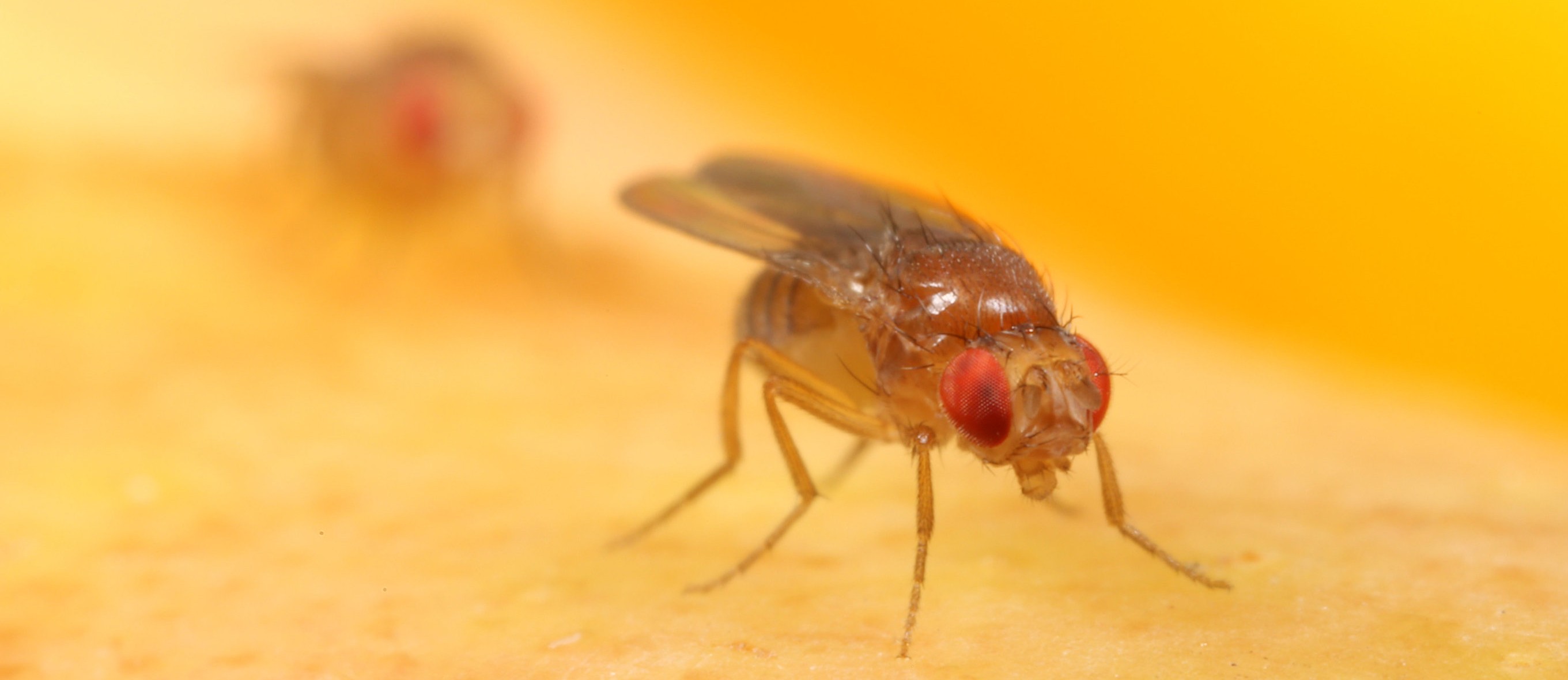
In our lab we see how different factors influence gene expression by measuring it in real time. We use Drosophila melanogaster (fruit flies) as our model organism, and engineer their DNA using CRISPR/Cas9 or PhiC31. Using the MS2-MCP system, we are able to visualize gene expression in a living fly. Repeating sequences of MS2 stem loops are inserted before a gene. As the gene is transcribed GFP binds to the MS2 stem loops and enables us to visualize the newly transcribed gene.

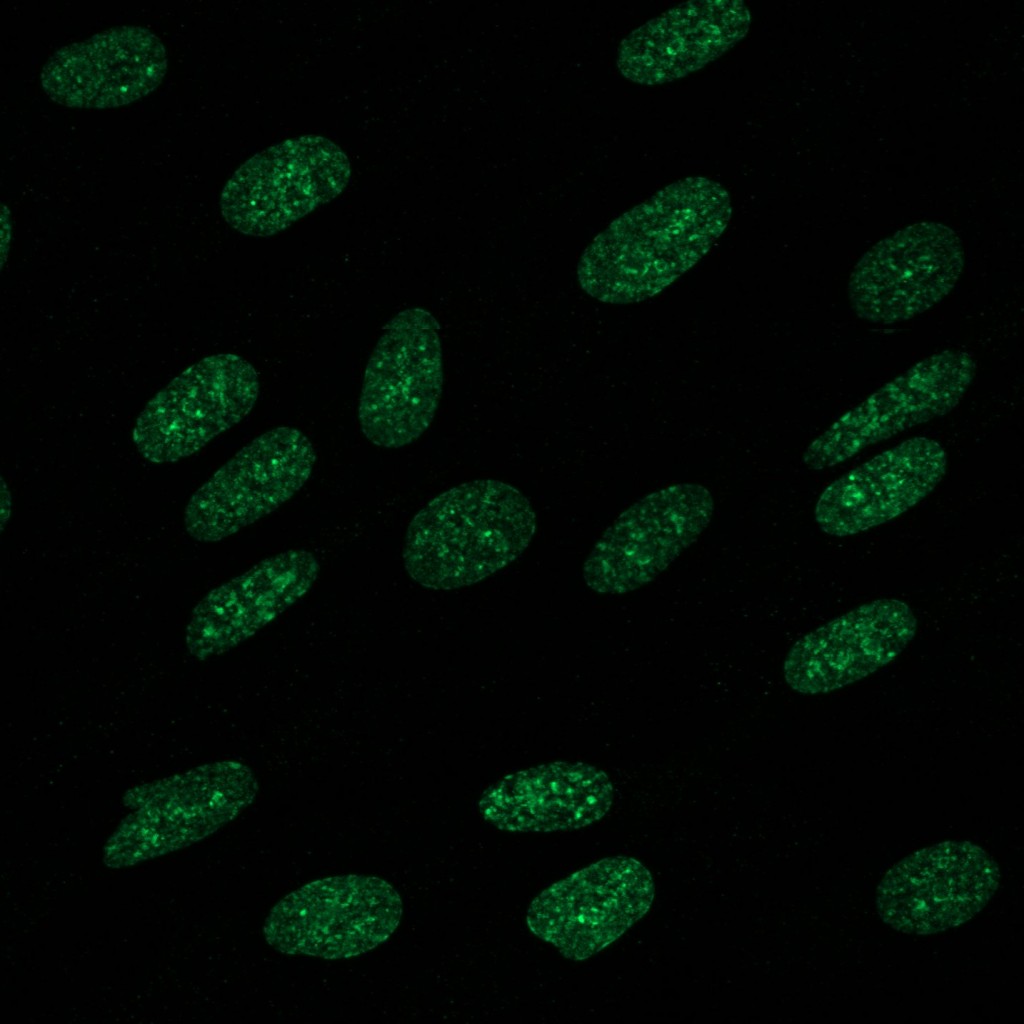
Epigenetics
Another method in which organisms influence gene expression is through epigenetic markers. By chemically modifying DNA, genes can be silenced or activated. A common epigenetic marker is the methylation of histones. Depending on where the methylation occurs, it can cause DNA to form tightly compacted heterochromatin, resulting in silencing the genes located within the compacted region. Using stem cells, we can use immunofluorescence to see certain methylation markers and measure the amount of heterochromatin present in a cell.